Watch our 2026 Commencement ceremony on Friday, May 29 at 1:30 p.m.!
Watch the Live Stream!
Former post-doc, Debjani Sihi, now an Assistant Professor at Emory University, led a paper published in Global Change Biology that presents a novel approach for representing complex soil microsite heterogeneity quantitatively using probability density functions of soil properties and microbial processes that affect these trace gas emissions.
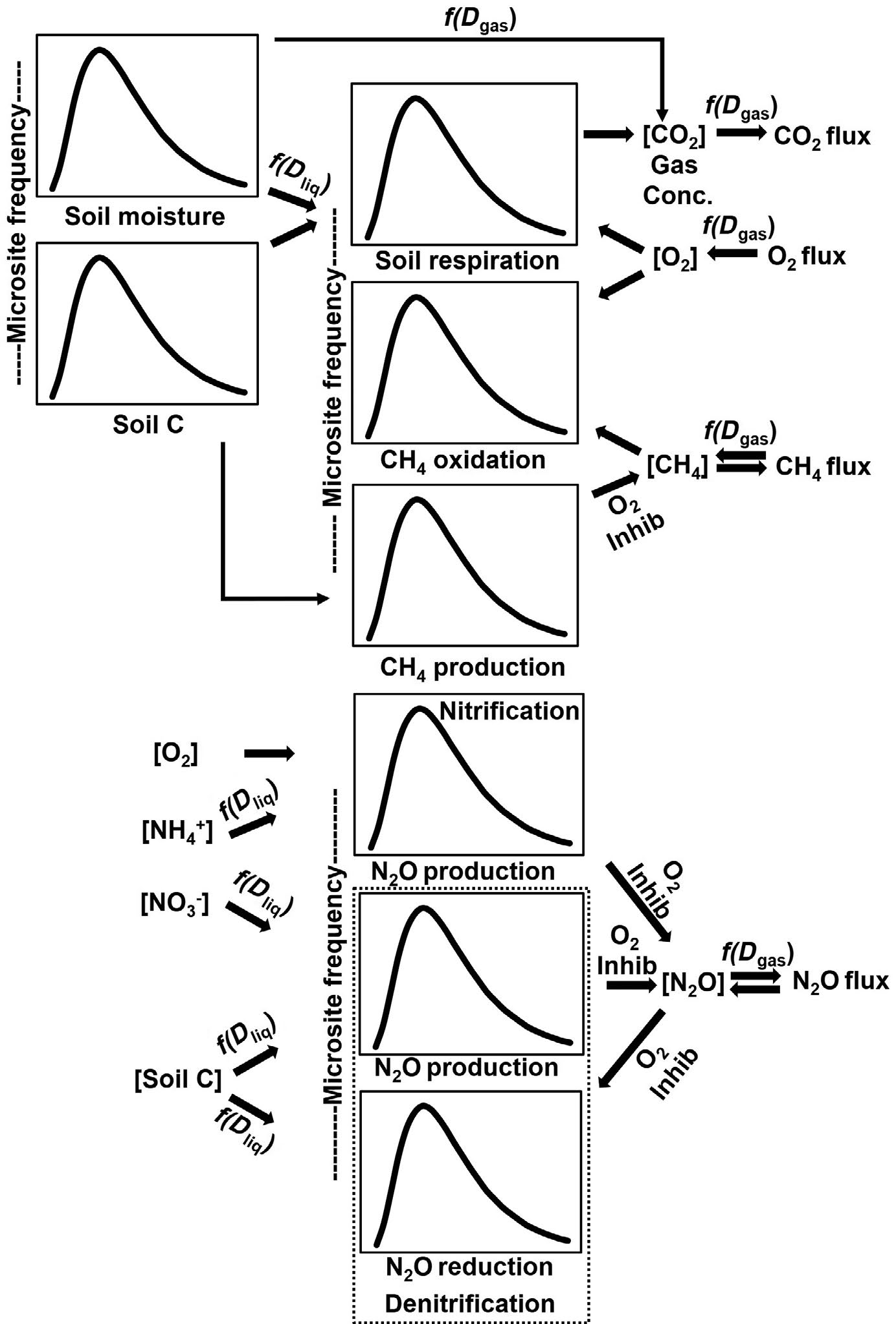
Rather than driving our model with mean measured soil carbon, water content, temperature, etc., we simulate probability distributions of these properties and use them to drive DAMM equations that calculate production and consumption of CO2, CH4, and N2O at each microsite. Aggregating the microsite simulations gives us flux simulations that are then compared to chamber measurements. While still computationally intense, our probability density function approach is computationally more efficient and viable than attempting to represent actual 3-D spatial heterogeneity of soil in a study plot. The results demonstrate how numerical representation of microsite heterogeneity can help us understand how a single soil profile can support both net uptake of CH4 (an oxidative process) and net consumption of N2O (a reductive process). While based on our understanding and theories of the controls on biological and physical soil processes, we use multiple constraints of three independent trace gas data streams simultaneously, thereby gaining confidence that the model is getting the right answer for the right reasons and properly simulating gas exchange, redox conditions, and microbial activity.